Tutorial: scikit-learn Pipelines
From orange juice 🍊 to food delivery 🛵
1 Introduction
Modern data analysis is much more than “fit a model.” Before a model can learn, we must understand the problem, prepare and clean the data, train, and evaluate it fairly and repeatably. The scikit-learn ecosystem in Python provides a unified API for all these steps.
In this assignment you will:
- explore how Pipelines describe and execute data-preparation steps,
- see how Pipelines bundle preprocessing and modeling together to prevent data leakage, and
- practice building, validating, and comparing models on two datasets.
We begin with a small orange juice dataset 🍊 to introduce scikit-learn Pipeline concepts clearly, then move on to a larger, more realistic food delivery dataset 🛵.
2 🎯 Learning objectives
By the end of this assignment, you will be able to:
- explain what
scikit-learnPipelines are and why they prevent leakage, - build preprocessing pipelines that handle categorical and numeric variables safely,
- combine preprocessing and modeling inside Pipelines,
- evaluate models with both random and time-aware cross-validation,
- tune and compare model types (e.g. LightGBM, SVM, regularized regression),
- engineer meaningful features such as distance and speed, and
- address class imbalance through undersampling and oversampling.
3 ⚙️ Set-up
3.1 Download dataset and notebook
You can download the dataset used in section 4 of this tutorial by clicking on the button below:
You can download the dataset used from section 5 of this tutorial by clicking on the button below:
Just like with all data in this course, place these new datasets into the data folder you created a few weeks ago.
You can also download the notebook that goes with this tutorial by clicking the button below:
3.2 Install needed libraries (if not done already)
conda install -c conda-forge scikit-learn lightgbm imbalanced-learn pandas numpy matplotlib seaborn3.3 Import needed libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import (
train_test_split, cross_validate,
KFold, TimeSeriesSplit, GridSearchCV, RandomizedSearchCV
)
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.impute import SimpleImputer
from sklearn.experimental import enable_iterative_imputer # noqa
from sklearn.impute import IterativeImputer
from sklearn.linear_model import LogisticRegression, ElasticNet
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor
from sklearn.metrics import (
f1_score, precision_score, recall_score, roc_auc_score,
classification_report, confusion_matrix, ConfusionMatrixDisplay,
mean_squared_error, mean_absolute_error, r2_score, root_mean_squared_error
)
import lightgbm as lgb
from lightgbm import LGBMClassifier, LGBMRegressor
from math import radians, cos, sin, asin, sqrt
from sklearn.calibration import calibration_curve4 🍊 Orange-juice warm-up
The aim of this section is to understand the logic of scikit-learn Pipelines through a compact, complete example. You will predict which brand of orange juice a customer buys—Citrus Hill (CH) or Minute Maid (MM)—using price, discounts, and loyalty information.
4.1 Load and explore the data
df_oj = pd.read_csv("data/OJ.csv")
print(df_oj.shape)
df_oj.head()(1070, 18)| Purchase | WeekofPurchase | StoreID | PriceCH | PriceMM | DiscCH | DiscMM | SpecialCH | SpecialMM | LoyalCH | SalePriceMM | SalePriceCH | PriceDiff | Store7 | PctDiscMM | PctDiscCH | ListPriceDiff | STORE | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | CH | 237 | 1 | 1.75 | 1.99 | 0.00 | 0.0 | 0 | 0 | 0.500000 | 1.99 | 1.75 | 0.24 | No | 0.000000 | 0.000000 | 0.24 | 1 |
| 1 | CH | 239 | 1 | 1.75 | 1.99 | 0.00 | 0.3 | 0 | 1 | 0.600000 | 1.69 | 1.75 | -0.06 | No | 0.150754 | 0.000000 | 0.24 | 1 |
| 2 | CH | 245 | 1 | 1.86 | 2.09 | 0.17 | 0.0 | 0 | 0 | 0.680000 | 2.09 | 1.69 | 0.40 | No | 0.000000 | 0.091398 | 0.23 | 1 |
| 3 | MM | 227 | 1 | 1.69 | 1.69 | 0.00 | 0.0 | 0 | 0 | 0.400000 | 1.69 | 1.69 | 0.00 | No | 0.000000 | 0.000000 | 0.00 | 1 |
| 4 | CH | 228 | 7 | 1.69 | 1.69 | 0.00 | 0.0 | 0 | 0 | 0.956535 | 1.69 | 1.69 | 0.00 | Yes | 0.000000 | 0.000000 | 0.00 | 0 |
Each row describes sales of orange juice for one week in one store. The outcome variable is Purchase, which records whether the customer bought CH or MM.
Other important variables include:
| Variable | Description |
|---|---|
PriceCH, PriceMM |
prices of Citrus Hill and Minute Maid |
DiscCH, DiscMM |
discounts for each brand |
LoyalCH |
loyalty toward Citrus Hill |
STORE, WeekofPurchase |
store ID and week number |
4.2 Sampling and stratified splitting
The full dataset has just over a thousand rows. To simulate a smaller study, we take a sample of 250 rows and then split it into training and test sets (60 % / 40 %). Because Purchase is categorical, we stratify so the CH/MM ratio remains constant.
Why 60/40 instead of 80/20? When the dataset is small, an 80/20 split leaves very few test examples, making metrics unstable. A 60/40 split provides enough data to train while still leaving a meaningful number of unseen samples for evaluation.
np.random.seed(123)
oj_sample = df_oj.sample(n=250, random_state=123)
oj_train, oj_test = train_test_split(
oj_sample, test_size=0.4, stratify=oj_sample["Purchase"], random_state=123
)
# check proportions
for name, subset in [("Full", df_oj), ("Sample 250", oj_sample),
("Train 60%", oj_train), ("Test 40%", oj_test)]:
counts = subset["Purchase"].value_counts(normalize=True)
print(f"{name}: CH={counts.get('CH', 0):.3f}, MM={counts.get('MM', 0):.3f}")Full: CH=0.610, MM=0.390
Sample 250: CH=0.584, MM=0.416
Train 60%: CH=0.587, MM=0.413
Test 40%: CH=0.580, MM=0.420Stratification ensures that both training and testing subsets keep the same mix of CH and MM. Without it, we could train mostly on one brand and evaluate on another—a subtle form of bias. Think of the dataset as a smoothie: 60 % orange 🍊 and 40 % mango 🥭. Every smaller glass should taste the same.
4.3 Feature engineering: add a season variable
WeekofPurchase records when the purchase occurred. Week numbers by themselves are not very informative, but they can reveal seasonal patterns—for instance, one brand might sell better in summer.
We’ll create a new column season based on week number. We also keep WeekofPurchase available as an identifier (it will be excluded from the model inside the Pipeline).
def add_season(df):
df = df.copy()
df["season"] = pd.cut(
df["WeekofPurchase"] % 52,
bins=[-1, 13, 26, 39, 52],
labels=["Winter", "Spring", "Summer", "Fall"]
).astype(str)
return df
oj_train = add_season(oj_train)
oj_test = add_season(oj_test)4.4 Understanding Pipeline
A scikit-learn Pipeline is a sequence of steps applied in order. It bundles all preprocessing transformers with a final estimator so the same transformations are applied identically during training, testing, and cross-validation.
Here is a complete Pipeline for our orange-juice problem:
numeric_features = ["PriceCH", "PriceMM", "DiscCH", "DiscMM", "LoyalCH", "STORE"]
categorical_features = ["season"]
# WeekofPurchase is intentionally excluded — acts as an ID
numeric_transformer = Pipeline([
("scaler", StandardScaler())
])
categorical_transformer = Pipeline([
("onehot", OneHotEncoder(handle_unknown="ignore", drop="first",
sparse_output=False))
])
preprocessor = ColumnTransformer([
("num", numeric_transformer, numeric_features),
("cat", categorical_transformer, categorical_features)
])
oj_pipeline = Pipeline([
("preprocessor", preprocessor),
("classifier", LogisticRegression(max_iter=1000, random_state=42))
])This block of code builds a complete, reproducible modelling workflow using scikit-learn’s Pipeline and ColumnTransformer.
Instead of manually:
- scaling numeric variables,
- encoding categorical variables,
- then fitting a model,
we combine everything into one coherent object.
4.4.1 Step 1 — Separate variables by type
numeric_features = ["PriceCH", "PriceMM", "DiscCH", "DiscMM", "LoyalCH", "STORE"]
categorical_features = ["season"]We explicitly tell the model:
- These variables are numeric → they need scaling.
- These variables are categorical → they need encoding.
WeekofPurchase is excluded because it behaves like an identifier rather than a predictive feature.
This separation is important because different types of variables require different preprocessing steps.
4.4.2 Step 2 — Define transformations for each type
4.4.2.1 Numeric variables
numeric_transformer = Pipeline([
("scaler", StandardScaler())
])We scale numeric variables so they:
- Have mean = 0
- Have standard deviation = 1
This is especially important for logistic regression because it:
- Improves numerical stability
- Makes coefficients comparable
- Helps regularisation behave properly
4.4.2.2 Categorical variables
categorical_transformer = Pipeline([
("onehot", OneHotEncoder(handle_unknown="ignore",
drop="first",
sparse_output=False))
])We use One-Hot Encoding to convert categories into binary indicators.
Key arguments:
handle_unknown="ignore"→ prevents errors if new categories appear in the test set.drop="first"→ avoids the dummy variable trap (perfect multicollinearity).sparse_output=False→ returns a regular NumPy array instead of a sparse matrix (easier to inspect).
4.4.3 Step 3 — Combine transformations with ColumnTransformer
preprocessor = ColumnTransformer([
("num", numeric_transformer, numeric_features),
("cat", categorical_transformer, categorical_features)
])ColumnTransformer applies:
- Scaling → only to numeric columns
- One-hot encoding → only to categorical columns
It ensures:
- No manual column slicing
- No risk of forgetting a transformation
- Clean separation of preprocessing logic
This is crucial for avoiding data leakage, because transformations will later be fitted only on the training data.
4.4.4 Step 4 — Build the full modelling pipeline
oj_pipeline = Pipeline([
("preprocessor", preprocessor),
("classifier", LogisticRegression(max_iter=1000, random_state=42))
])This creates one unified object that:
- Applies preprocessing
- Then fits a logistic regression model
When you run:
oj_pipeline.fit(X_train, y_train)scikit-learn automatically:
- Fits the scaler on the training data
- Fits the encoder on the training data
- Transforms the training data
- Trains the logistic regression model
And when you run:
oj_pipeline.predict(X_test)it automatically:
- Applies the same fitted transformations to the test set
- Then makes predictions
No manual intervention needed.
If you had scaled the data before splitting into training and test sets, the scaler would learn the mean and variance from the entire dataset — including the test set.
That would leak information from the test set into the model.
Because the scaler lives inside the Pipeline, it is fitted only when .fit() is called on the training data.
This guarantees honest evaluation.
After preprocessing:
- Numeric variables are scaled.
- The categorical variable
seasonbecomes multiple binary columns.
For example, if season has values:
Fall, Winter, Spring, SummerOne-hot encoding (with drop="first") produces:
season_Spring
season_Summer
season_WinterEach row now contains only numbers.
The logistic regression model works purely on this transformed numeric matrix.
4.5 Why is this better than doing it step-by-step?
Without a Pipeline, you would need to:
- Fit the scaler separately
- Transform training data
- Transform test data
- Fit the encoder
- Keep track of column order
- Manually pass transformed arrays into the model
That is:
- Error-prone
- Harder to reproduce
- Easier to introduce leakage
A Pipeline guarantees:
- Correct order of operations
- Consistent preprocessing at train and test time
- Cleaner code
- Easier cross-validation
4.5.1 Conceptual Summary
Think of a Pipeline as:
A recipe that defines how raw data becomes predictions.
It encodes the entire modelling logic into a single, reusable object.
Raw data (mixed numeric + categorical)
↓
ColumnTransformer
• Scale numeric features
• One-hot encode categorical features
↓
Fully numeric feature matrix
↓
Logistic Regression
↓
Predictions
The classifier never sees raw data — it only sees the transformed numeric matrix.
- “Where are my transformed columns?” The Pipeline hides intermediate outputs. You can inspect them using:
oj_pipeline.named_steps["preprocessor"].get_feature_names_out()“Why do I see more coefficients than original variables?” Because one categorical variable can expand into multiple binary features.
“Why is scaling necessary?” Logistic regression with regularisation is sensitive to feature scale.
One major advantage of Pipelines is that they work seamlessly with cross-validation:
cross_val_score(oj_pipeline, X, y, cv=5)Each fold:
- Refits the scaler
- Refits the encoder
- Refits the classifier
All automatically.
This makes Pipelines essential for proper model evaluation.
4.5.2 StandardScaler
StandardScaler() transforms each numeric variable by:
- Subtracting its mean
- Dividing by its standard deviation
After scaling, each variable has:
- Mean = 0
- Standard deviation = 1
This process is often called standardisation or z-score scaling.
4.5.2.1 Why standardise?
Many machine learning models are sensitive to the scale of the input variables.
For example:
- Logistic regression (with regularisation) penalises large coefficients — if variables are on very different scales, the penalty behaves unevenly.
- Support Vector Machines (SVMs) depend on distances in feature space.
- K-Nearest Neighbours (KNN) is entirely distance-based.
If one variable ranges between 0–1 and another ranges between 0–10,000, the larger-scale variable can dominate the model simply because of its units.
Standardising puts all numeric predictors on a comparable scale and:
- Improves numerical stability
- Makes regularisation behave properly
- Helps optimisation converge more reliably
- Makes coefficients more interpretable in relative terms
4.6 Choose a model
We will use logistic regression as a baseline model.
Why start here?
- It is a strong and widely used benchmark for binary classification.
- It is fast to train and easy to interpret.
- With regularisation (the default in
scikit-learn), it provides a sensible bias–variance trade-off. - If a simple linear model performs well, more complex models may not be necessary.
In modelling, starting simple and increasing complexity only if needed is good practice.
4.7 Build and fit the Pipeline
A Pipeline combines preprocessing and modeling so the correct steps are always applied in the correct order. Crucially, all preprocessing statistics (means, standard deviations, encoder levels) are learned only from the training data when you call fit().
X_train = oj_train[numeric_features + categorical_features]
y_train = oj_train["Purchase"]
X_test = oj_test[numeric_features + categorical_features]
y_test = oj_test["Purchase"]
oj_pipeline.fit(X_train, y_train)4.8 Fit and evaluate the model
We now evaluate the model on the held-out test set, which was not used during training.
y_pred = oj_pipeline.predict(X_test)
class_index = list(oj_pipeline.classes_).index("CH")
y_proba = oj_pipeline.predict_proba(X_test)[:, class_index]
print("F1: ", round(f1_score(y_test, y_pred, pos_label="CH"), 3))
print("Precision:", round(precision_score(y_test, y_pred, pos_label="CH"), 3))
print("Recall: ", round(recall_score(y_test, y_pred, pos_label="CH"), 3))
print("ROC AUC: ", round(roc_auc_score((y_test == "CH").astype(int), y_proba), 3))We explicitly select the probability corresponding to class "CH" to ensure consistency with the chosen positive label.
F1: 0.914
Precision: 0.914
Recall: 0.914
ROC AUC: 0.956The high F₁ and ROC AUC suggest that even a simple baseline model separates the two classes effectively.
4.8.1 How fit() and predict() work
When you call fit() on a Pipeline, scikit-learn first fits each transformer on the training data: it computes the statistics required for preprocessing—means and standard deviations for scaling, the set of levels for categorical variables, etc.—and stores them inside the Pipeline. It then fits the model (here, a logistic regression) to the processed training data.
Later, predict() re-applies the trained transformers to the test data using those same stored statistics. No new averages or encodings are computed; the test data remain unseen during training. This is what prevents data leakage—the model never peeks at future information.
4.8.2 Choosing appropriate metrics
Because the classes are not perfectly balanced, accuracy can be misleading. A model that always predicts the majority class might look “accurate” but tell us little. Instead we evaluate:
- Precision: when the model predicts
CH, how often is it correct? - Recall: of all actual
CHpurchases, how many did it find? - F₁-score: the harmonic mean of precision and recall, balancing both.
- ROC AUC: the area under the ROC curve, summarizing discrimination across thresholds.
These measures together give a more complete picture of performance.
- F₁ reflects performance at the default 0.5 classification threshold.
- ROC AUC evaluates ranking quality across all possible thresholds.
Looking at both helps distinguish threshold-specific performance from overall discrimination ability.
- Replace
LogisticRegression()with another classifier you know, such asRandomForestClassifier()orSVC(kernel="rbf", probability=True). - Rerun the Pipeline and evaluation.
- Compare F₁ and ROC AUC. Which model performs better, and why might that be?
5 Food delivery dataset 🛵
You are now ready to apply the same ideas to a realistic dataset.
We will use a public Food Delivery dataset from Kaggle containing restaurant, delivery-person, and environmental details along with the delivery time in minutes (we’re using the dataset called train on Kaggle).
Your objectives are to:
- Predict delivery time (
Time_taken(min)) using regression. - Compare cross-sectional and time-aware validation.
- Later, classify deliveries by speed.
5.1 Load and explore the data
food = pd.read_csv(
"data/food-delivery.csv",
na_values=["", "NA", "NaN", "null"],
skipinitialspace=True
)
numeric_cols = [
"Delivery_person_Age",
"Delivery_person_Ratings",
"multiple_deliveries"
]
for col in numeric_cols:
food[col] = pd.to_numeric(food[col], errors="coerce")
food["Order_Date"] = pd.to_datetime(food["Order_Date"], dayfirst=True)
food["Order_Week"] = food["Order_Date"].dt.isocalendar().week.astype(int)
food = food.sort_values("Order_Date").reset_index(drop=True)
print(food.shape)
food.info()Real-world datasets often contain placeholder values with hidden whitespace (e.g. "NaN "). These are not automatically treated as missing in pandas.
Always strip whitespace before numeric conversion to ensure missing values are properly encoded.
The dataset includes restaurant and delivery coordinates, information about the delivery person, weather, traffic, and more.
We first convert Order_Date from a string into a proper datetime object using:
pd.to_datetime(df["Order_Date"], dayfirst=True)5.1.0.1 Why dayfirst=True?
Dates such as "03-04-2023" are ambiguous:
- Without
dayfirst=True, pandas may interpret this as March 4th. - With
dayfirst=True, it is interpreted as 3 April.
Because the dataset uses a day–month–year format, we must explicitly specify this to avoid silent misinterpretation. If we do not, the dates may be parsed incorrectly without raising an error — which would distort any time-based analysis.
After converting to datetime, we create Order_Week (the ISO week number) to group all orders from the same week.
5.1.0.2 Why group by week rather than day or month?
- Daily data can be too granular: some days may contain only a small number of deliveries, leading to noisy estimates.
- Monthly data are often too coarse and may hide short-term patterns such as weather shocks or traffic disruptions.
- Weekly aggregation provides a balance: enough observations per period for stability, while still capturing meaningful temporal variation.
This choice will also matter later when we implement time-aware validation.
The variable Time_taken(min) is stored as text values such as "(min) 21". Convert it to numeric values like 21 using a suitable string operation and transformation.
💡 Question: Why do you think this conversion is important before modeling?
5.2 Create a seasonal feature
We’ll create a new column season based on week number.
def assign_season(week):
if week in range(1, 14):
return "Winter"
elif week in range(14, 27):
return "Spring"
elif week in range(27, 40):
return "Summer"
else:
return "Autumn"
food["Season"] = food["Order_Week"].apply(assign_season)5.3 Compute a simple distance feature
Delivery time should depend strongly on distance. We can approximate distance using the Haversine formula, which computes the great-circle (shortest) distance between two points on Earth.
# Earth radius in kilometres
R = 6371
# Convert latitude and longitude from degrees to radians
# (Trigonometric functions in NumPy expect radians)
lat1 = np.radians(food["Restaurant_latitude"].to_numpy())
lon1 = np.radians(food["Restaurant_longitude"].to_numpy())
lat2 = np.radians(food["Delivery_location_latitude"].to_numpy())
lon2 = np.radians(food["Delivery_location_longitude"].to_numpy())
# Differences in coordinates
dlat = lat2 - lat1
dlon = lon2 - lon1
# Haversine formula (vectorised)
food["distance_km"] = 2 * R * np.arcsin(
np.sqrt(
np.sin(dlat / 2) ** 2 +
np.cos(lat1) * np.cos(lat2) * np.sin(dlon / 2) ** 2
)
)Even a rough, straight-line distance captures a large share of the variation in delivery times. It reflects the physical effort of travel even when we do not model exact routes, traffic lights, or congestion patterns.
In many real-world problems, a simple geometric feature like this can dramatically improve predictive performance.
We compute great-circle distance using the haversine formula, which measures the shortest path between two points on a sphere.
Key implementation details:
- Trigonometric functions in NumPy expect radians, not degrees — hence the use of
np.radians(). - The formula is applied to entire arrays at once, not row by row.
- This is called vectorisation: NumPy performs the computation in compiled C code, making it much faster than
.apply(axis=1).
Unlike pairwise distance functions (e.g sklearn.metrics.pairwise.haversine_distances()), this implementation computes distances elementwise (row by row conceptually, but efficiently under the hood), so it scales linearly with the dataset size (the results are reasonable memory usage and reasonable running time!).
Earlier we could have used a pairwise distance function (e.g sklearn.metrics.pairwise.haversine_distances()), but that would compute a full \(n \times n\) distance matrix.
With 45,000+ rows, that would require billions of distance calculations and several gigabytes of memory.
The vectorised NumPy implementation instead computes only the distances we actually need — one per row — making it both faster and memory-efficient.
Each distance_km value depends only on its own row (restaurant and delivery coordinates).
No averages, global statistics, or shared information are used. Therefore, creating this feature before the train/test split does not introduce data leakage.
5.4 Create time-of-day features
Delivery times don’t depend solely on distance or traffic — they also vary throughout the day. Morning commutes, lunch peaks, and late-night orders each impose different operational pressures on both restaurants and delivery drivers.
We’ll capture these daily patterns by creating new variables describing the time of order placement and time of pickup, both as numeric hours and as interpretable day-part categories.
# ------------------------------------------------------------------
# 1️⃣ Clean invalid time entries
# ------------------------------------------------------------------
food["Time_Orderd"] = food["Time_Orderd"].replace(
["NaN", "", "null", "0"], np.nan
)
food["Time_Order_picked"] = food["Time_Order_picked"].replace(
["NaN", "", "null", "0"], np.nan
)
# ------------------------------------------------------------------
# 2️⃣ Parse clock times into datetime objects
# ------------------------------------------------------------------
food["Order_dt"] = pd.to_datetime(
food["Time_Orderd"],
format="%H:%M:%S",
errors="coerce"
)
food["Pickup_dt"] = pd.to_datetime(
food["Time_Order_picked"],
format="%H:%M:%S",
errors="coerce"
)
# ------------------------------------------------------------------
# 3️⃣ Extract hour of day (0–23)
# ------------------------------------------------------------------
food["Order_Hour"] = food["Order_dt"].dt.hour
food["Pickup_Hour"] = food["Pickup_dt"].dt.hour
# ------------------------------------------------------------------
# 4️⃣ Create coarse time-of-day categories
# ------------------------------------------------------------------
conditions = [
food["Order_Hour"].between(5, 11),
food["Order_Hour"].between(12, 16),
food["Order_Hour"].between(17, 21),
]
choices = ["Morning", "Afternoon", "Evening"]
food["Order_Time_of_Day"] = np.select(
conditions,
choices,
default="Night"
)
conditions_pickup = [
food["Pickup_Hour"].between(5, 11),
food["Pickup_Hour"].between(12, 16),
food["Pickup_Hour"].between(17, 21),
]
food["Time_of_Day"] = np.select(
conditions_pickup,
["Morning", "Afternoon", "Evening"],
default="Night"
)⚠️ Some entries in Time_Orderd are invalid (e.g., "NaN", "", or "0") and can’t be converted to clock times. To avoid errors, we explicitly replace these with NaN before parsing. Missing times are later handled inside the Pipeline’s imputation step.
This block performs four distinct transformations:
1️⃣ Clean invalid entries
Some rows contain placeholders such as "NaN", "", "null", or "0". These are not valid clock times. We explicitly convert them to NaN so that pandas recognises them as missing values.
2️⃣ Parse strings into proper time objects
We convert text values like "13:42:00" into datetime objects using:
format="%H:%M:%S"→ ensures consistent and fast parsingerrors="coerce"→ invalid values becomeNaT(missing time)
This step makes time information machine-readable.
3️⃣ Extract the hour (0–23)
Using .dt.hour, we derive numeric features:
Order_HourPickup_Hour
These capture daily demand and traffic cycles in a simple, interpretable form.
4️⃣ Create broader day-part categories
Using np.select(), we group hours into:
- Morning (5–11)
- Afternoon (12–16)
- Evening (17–21)
- Night (default)
This creates coarser behavioural signals that may capture nonlinear effects (e.g., dinner rush vs. late-night deliveries).
Importantly, all operations are fully vectorised — no row-wise loops — making the transformation efficient even for large datasets.
Missing values created here will later be handled by the Pipeline’s imputation step.
When deriving categorical summaries from numeric variables (like converting Order_Hour to Order_Time_of_Day), you must explicitly preserve missingness.
The function above returns np.nan when Order_Hour is missing, so that missing order timestamps propagate correctly into the derived columns — otherwise, the model might incorrectly assume those rows are "Night" orders!
5.4.1 🌆 Why this matters
Raw timestamps like “13:42:00” are not directly meaningful for machine learning models:
- They are too granular
- They are cyclical (23:59 is close to 00:01)
- Their numeric representation has no linear meaning
By extracting:
- Hour (0–23) → captures daily demand and traffic cycles
- Day-part categories → captures broader behavioural patterns
we convert raw time strings into structured, interpretable features.
This transformation turns clock data into operational signals about:
- Restaurant load
- Traffic conditions
- Customer demand cycles
| Feature | Type | Operational meaning | Expected effect on delivery time |
|---|---|---|---|
Order_Hour |
Numeric (0–23) | The exact hour when the customer placed the order | Reflects demand cycles — lunch (12–14h) and dinner (19–21h) peaks increase restaurant load and prep time. |
Pickup_Hour |
Numeric (0–23) | The exact hour when the courier picked up the order | Reflects road and traffic conditions — peak commute hours (8–10h, 17–20h) slow travel. |
Order_Time_of_Day |
Categorical (Morning/Afternoon/Evening/Night) | Coarse summary of restaurant-side demand | Morning orders (e.g., breakfast deliveries) tend to be faster; dinner rushes slower. |
Time_of_Day |
Categorical (Morning/Afternoon/Evening/Night) | Coarse summary of driver-side conditions | Evening and night deliveries face heavier congestion and longer travel times. |
5.5 Create a cross-sectional dataset and training/test split
So far, we have engineered features using the full dataset to understand its structure and missingness patterns.
We now switch perspective: instead of modelling the entire time span, we isolate a single, dense operational period — the busiest week i.e the week with most orders— and treat it as a cross-section.
This allows us to study model behaviour in a stable operational context, without temporal drift.
busiest_week = food["Order_Week"].value_counts().idxmax()
cross_df = food[food["Order_Week"] == busiest_week].copy()
print(f"Busiest week: {busiest_week}, rows: {len(cross_df)}")Busiest week: 10, rows: 7554Why the busiest week? It provides enough examples for both training and validation within a consistent operational context (similar traffic, weather, and demand). This design isolates one dense period to study model behavior without temporal drift.
Split the cross_df data into training and test sets. Use about 80 % of the data for training and 20 % for testing. Set a random seed for reproducibility.
5.6 Handle missing data with iterative imputation
Only a few predictors contain missing data:
Delivery_person_RatingsDelivery_person_Agemultiple_deliveries
Rather than filling these with simple averages or modes, we will impute them using iterative (bagged-tree) imputation.
5.6.1 What is iterative imputation?
scikit-learn’s IterativeImputer implements multivariate imputation by modelling each incomplete variable as a function of the others:
- For each variable with missing values, an estimator is trained on the complete rows using all other variables as features.
- Missing entries are predicted from that model.
- The process cycles through all variables with missingness until convergence.
The process cycles through variables multiple times, updating predictions until values stabilise (convergence).
The estimator we use in conjunction with IterativeImputer is a Random Forest regressor (RandomForestRegressor as the base estimator). This choice allows us to capture nonlinear relationships and interactions between features during imputation, which can lead to more accurate estimates than simple mean or median imputation.
Before you decide how to impute, always explore where data are missing.
- Use
df.isnull().sum()ordf.describe()to inspect missing values in each column. - Identify which variables have the most missingness.
- Write two or three sentences: do you think the missingness is random or related to certain conditions (e.g., specific cities, traffic levels, or festivals)?
5.7 Build a regression Pipeline
You will now build a `scikit-learn Pipeline to predict Time_taken(min) from operational predictors.
Do not hard-code transformations outside the Pipeline. All preprocessing must live inside it.
Your Pipeline should:
- Handle missing values appropriately.
- Encode categorical predictors.
- Scale numeric predictors.
- Be compatible with cross-validation.
- Avoid data leakage.
You may consult:
- The
scikit-learndocumentation on Pipelines and ColumnTransformer: https://scikit-learn.org/stable/modules/compose.html - The Orange Juice example at the start of this notebook, where we constructed a full preprocessing + modelling Pipeline.
Think carefully about:
- Which variables are meaningful predictors.
- Which variables are identifiers or raw fields that should not be used.
- Whether some variables have no variation in a cross-sectional setting.
- Whether engineered variables already replace raw ones.
You are responsible for defining:
numeric_features = [...]
categorical_features = [...]and constructing an appropriate ColumnTransformer.
A Pipeline must be:
- Defined before splitting (that’s fine),
- But fitted only on training data.
You will later integrate this Pipeline into cross-validation. If preprocessing is done outside the Pipeline, your results are invalid.
5.8 Choose and specify a regression model – LightGBM
We will use LightGBM, a modern gradient-boosted-tree algorithm that is very efficient on large tabular data.
Both LightGBM and XGBoost implement gradient boosting: a sequence of small trees is trained, each correcting residual errors from previous trees. The difference lies in how they grow trees and manage splits:
- XGBoost grows trees level-wise (balanced): it splits all leaves at one depth before going deeper. This produces symmetric trees and is often robust on small or moderately sized data.
- LightGBM grows trees leaf-wise: it always expands the single leaf that most reduces the loss, yielding deeper, targeted trees. Together with histogram-based binning of features, this makes LightGBM faster and often stronger on large, sparse, or high-cardinality data.
5.8.1 Example specification
lgb_reg = LGBMRegressor(
n_estimators=500,
learning_rate=0.1,
max_depth=6,
min_child_samples=10, # slightly larger
verbose=-1, # suppress warnings
random_state=42
)5.8.2 Common LightGBM tuning parameters
| Parameter | What it controls | Intuition |
|---|---|---|
n_estimators |
total number of trees | more trees can capture more structure but risk overfitting unless learning_rate is small |
learning_rate |
step size for each tree’s contribution | small = slower but safer learning; large = faster but riskier |
max_depth |
maximum depth of a single tree | deeper trees capture complex interactions but can overfit |
min_child_samples |
minimum observations in a leaf | prevents tiny leaves that memorize noise |
num_leaves |
number of leaves per tree | key LightGBM parameter; larger = more complex trees |
colsample_bytree |
fraction of predictors sampled at each split | adds randomness, reduces correlation among trees |
When might you prefer XGBoost? On smaller datasets, when stability and interpretability matter. When might you prefer LightGBM? For speed on larger tables, many categorical dummies, or limited tuning time.
5.9 Combine preprocessor and model inside a Pipeline
Once your ColumnTransformer preprocessor and model are ready, combine them in a single Pipeline so preprocessing and modeling stay linked. The Pipeline automatically applies preprocessing before fitting or predicting.
reg_pipeline = Pipeline([
("preprocessor", preprocessor),
("regressor", lgb_reg)
])5.10 Evaluate the cross-sectional model
Fit your Pipeline on the training set and evaluate predictions on the test set. Use regression metrics such as RMSE, MAE, and R².
Compute and report these metrics for your model. How accurate is the model on unseen data? Does it tend to over- or under-predict delivery time?
Numerical metrics tell part of the story — but to see how your model behaves, you need to inspect residuals (the differences between actual and predicted delivery times).
Try plotting:
Predicted vs. actual delivery times → Are most points near the diagonal (perfect predictions)? → Do errors grow with longer delivery times?
Residuals vs. predicted values → Are residuals centered around zero (no systematic bias)? → Do you see patterns that might suggest missing predictors?
Residuals by time of day → Are certain times (e.g., evening rush hours) consistently over- or under-predicted?
Write a short paragraph summarizing what you notice. Does the model perform equally well for all deliveries, or does it struggle under specific conditions?
5.10.1 Cross-validation on the cross-sectional data
So far, you evaluated the model with a single train/test split. In a cross-sectional setting (no time ordering), it is common to use k-fold cross-validation, which splits the training data into k random folds, training on k – 1 folds and validating on the remaining fold in turn. This yields a more robust estimate of performance stability.
Run a 5-fold cross-validation on the cross-sectional training data. Compare the average RMSE and R² to those from the single test-set evaluation. Are they similar? Which estimate do you trust more?
Use the scikit-learn documentation if needed: https://scikit-learn.org/stable/modules/model_evaluation.html
When using cross-validation, some error-based metrics (e.g., RMSE, MAE) may be returned as negative values.
This is because scikit-learn uses a “higher is better” convention for model selection.
Make sure you interpret these values correctly when reporting results.
5.11 Time-aware validation with TimeSeriesSplit
In time-dependent data, future information must not influence the model’s training. To mimic real-world deployment, we roll forward through time: train on past weeks, validate on the next week(s).
# Chronological split: 80% train, 20% test
cutoff = int(len(food) * 0.8)
long_train = food.iloc[:cutoff].copy()
long_test = food.iloc[cutoff:].copy()
# Time-aware CV: expanding windows
cv_long = TimeSeriesSplit(n_splits=6)Never define cross-validation folds on the entire dataset. If you did, the model could “peek” at future data while training, inflating performance. Instead, use only the training portion of your longitudinal split — here called long_train.
5.11.1 How TimeSeriesSplit works
TimeSeriesSplit creates a sequence of rolling resamples ordered by row index. Because the data were sorted by Order_Date before splitting, row order corresponds to time order. Each successive fold uses all past rows as training data and the next block of rows as validation — exactly like an expanding window. Unlike KFold, TimeSeriesSplit never shuffles the data and never allows future observations to enter the training set. Each fold expands the training window forward in time.
for i, (train_idx, val_idx) in enumerate(cv_long.split(long_train)):
train_dates = long_train.iloc[train_idx]["Order_Date"]
val_dates = long_train.iloc[val_idx]["Order_Date"]
print(f"Fold {i+1}: train {train_dates.min().date()} → {train_dates.max().date()}"
f" | val {val_dates.min().date()} → {val_dates.max().date()}")Fold 1: train 2022-02-11 → 2022-02-16 | val 2022-02-16 → 2022-03-03
Fold 2: train 2022-02-11 → 2022-03-03 | val 2022-03-03 → 2022-03-08
Fold 3: train 2022-02-11 → 2022-03-08 | val 2022-03-08 → 2022-03-13
Fold 4: train 2022-02-11 → 2022-03-13 | val 2022-03-13 → 2022-03-18
Fold 5: train 2022-02-11 → 2022-03-18 | val 2022-03-18 → 2022-03-24
Fold 6: train 2022-02-11 → 2022-03-24 | val 2022-03-24 → 2022-03-29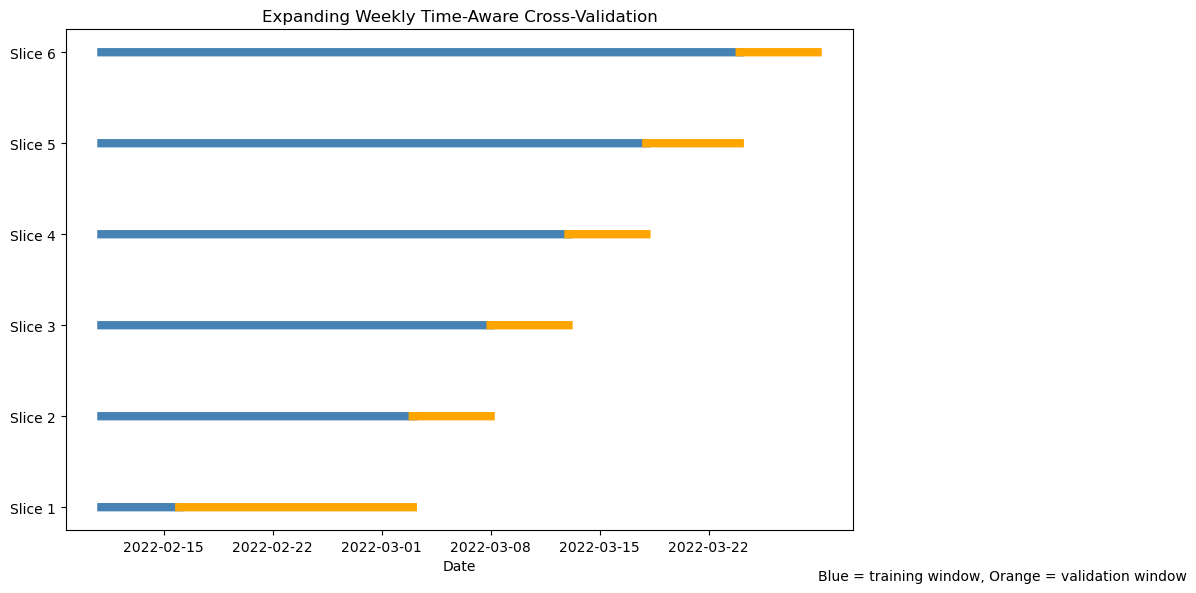
Visualising the folds (code)
# Ensure chronological order
long_train = long_train.sort_values("Order_Date").reset_index(drop=True)
dates = long_train["Order_Date"]
fig, ax = plt.subplots(figsize=(12, 6))
for i, (train_idx, val_idx) in enumerate(cv_long.split(long_train)):
train_dates = dates.iloc[train_idx]
val_dates = dates.iloc[val_idx]
# Training window
ax.plot(
train_dates,
[i] * len(train_dates),
color="steelblue",
linewidth=6
)
# Validation window
ax.plot(
val_dates,
[i] * len(val_dates),
color="orange",
linewidth=6
)
ax.set_yticks(range(i + 1))
ax.set_yticklabels([f"Slice {j+1}" for j in range(i + 1)])
ax.set_xlabel("Date")
ax.set_title("Expanding Weekly Time-Aware Cross-Validation")
ax.text(
dates.iloc[-1],
-0.8,
"Blue = training window, Orange = validation window"
)
plt.tight_layout()
plt.show()5.12 Pipeline and modeling for longitudinal validation
Repeat the full process—Pipeline, model, and evaluation—on the longitudinal setup. Then train and evaluate the model using the time-aware splits created above.
Use cross_validate() with cv=cv_long to fit your LightGBM Pipeline on long_train. Collect metrics (RMSE, MAE, R²) and summarize them.
5.13 Challenge: comparing designs
Compare your cross-sectional and longitudinal results.
- Are the results directly comparable? Why or why not?
- Which design better represents real-world performance, and why?
- How does time-aware validation change your interpretation of model quality?
5.14 🧗 Challenge: try another model
Repeat the workflow using a different algorithm of your choice (for example, Elastic-Net, SVM, or KNN). Do not copy specifications here — consult lectures, 📚 labs W02–W05, and the reading week homework for syntax, recommended preprocessing, and tuning guidance.
Evaluate the alternative model on both cross-sectional and time-aware folds and compare results.
- Which model generalized best over time?
- Did any model appear acceptable under random folds but degrade under time-aware validation?
- Which preprocessing steps (such as normalization) were essential for your chosen model, and why?
- Would your preferred model still perform well if new conditions appeared (e.g., “Summer” deliveries or new cities)?
6 Classifying deliveries by speed 🚴♀️
So far, you have treated delivery time as a continuous variable and built regression models to predict it.
We will now reframe the same problem as a classification task: predicting whether a delivery is fast, medium, or slow based on its characteristics.
This shift lets you practice classification workflows and also experiment with techniques for handling class imbalance while keeping the same modelling discipline: time-aware validation, leakage prevention, and Pipelines. .
6.1 Define delivery-speed classes
Rather than using arbitrary thresholds (for example “below 30 minutes = fast”), we can define speed relative to distance.
A fairer comparison is delivery speed = distance / time in km per minute (or per hour).
food["speed_km_min"] = food["distance_km"] / food["Time_taken(min)"]
q33 = food["speed_km_min"].quantile(0.33)
q67 = food["speed_km_min"].quantile(0.67)
def assign_speed_class(s):
if s >= q67:
return "Fast"
elif s <= q33:
return "Slow"
else:
return "Medium"
food["speed_class"] = food["speed_km_min"].apply(assign_speed_class)
food["speed_class"] = pd.Categorical(food["speed_class"],
categories=["Slow", "Medium", "Fast"],
ordered=True)This approach adapts automatically to the dataset’s scale and ensures that roughly one-third of deliveries fall in each class. You can adjust the quantile cut-offs if you wish to make the “Fast” or “Slow” categories narrower.
If you prefer a binary classification (for example fast vs not fast), combine “Slow” and “Medium” into a single category. Think about which framing would be more relevant for an operations team.
6.2 Explore class balance
Before training any classifier, you should always examine how balanced your outcome variable is.
Understanding class proportions helps you decide:
- Whether accuracy is meaningful,
- Whether macro- or weighted-averaged metrics are appropriate,
- Whether class weighting or resampling might be needed.
- Compute the count of observations in each
speed_class. - Compute the proportion of each class.
- Present your results in a clear table.
- Briefly comment on whether the dataset appears balanced.
Based on your findings, decide whether macro-averaged metrics are appropriate.
6.3 🤺 Your turn: build a classification Pipeline
Split the data into training/test sets and create a Pipeline that predicts speed_class from the relevant predictors (distance, traffic, weather, vehicle type, etc.).
Follow the same logic as before but adapt it for classification:
- Impute missing numeric values (tree-based imputation is fine, but keep it efficient).
- Impute missing categorical values with
SimpleImputer(strategy="most_frequent"). - One-hot encode categorical variables with
OneHotEncoder(handle_unknown="ignore"). - Normalize numeric predictors with
StandardScaler. - Optionally explore resampling using
imblearnonly in random CV, not time-aware CV.
Consult the scikit-learn ColumnTransformer documentation for details.
You may see these tools in imbalanced-learn:
from imblearn.pipeline import Pipeline as ImbPipeline
from imblearn.over_sampling import RandomOverSampler
from imblearn.under_sampling import RandomUnderSamplerWhat they are for:
ImbPipelineallows resampling steps to live inside a Pipeline, so resampling happens correctly within each training fold during cross-validation.RandomOverSamplerduplicates minority-class examples in the training fold.RandomUnderSamplerdiscards some majority-class examples in the training fold.
Important (design caveat): these assume that rows can be shuffled freely. So they are suitable for random cross-validation (cross-sectional design), but not for time-aware CV such as TimeSeriesSplit.
In this section’s solution, we tune with time-aware CV, so we do not use oversampling/undersampling here. We mention them to guide further exploration later.
⚠️ Important:
Upsampling and downsampling assume that observations can be freely shuffled between resamples. They are suitable for random cross-validation, where all rows are exchangeable. However, they should not be used inside time-aware cross-validation such as TimeSeriesSplit. Doing so would destroy the chronological structure and introduce future information into past folds, leading to data leakage. If you wish to handle imbalance in time-series data, use class-weighting (see this page and this page for more explanations) or use metrics that account for imbalance (macro-F1).
6.4 Choose a classification model
You will now build a classifier to predict speed_class.
You may choose any model covered in lectures (e.g., logistic regression, SVM, KNN, random forest, gradient boosting).
Consult:
- Your lecture notes
- Labs W02–W05
- The Orange Juice example at the beginning of this notebook
Your classifier must:
- Be combined with your preprocessing object inside a
Pipeline. - Produce class probabilities (required later for ROC AUC and calibration).
- Select a classifier.
- Combine it with your preprocessing step inside a
Pipeline. - Briefly justify your choice of model.
- Consider whether unequal error costs might matter in deployment.
6.5 Time-aware tuning
Because this is a chronological prediction problem, model validation must respect time ordering.
We therefore tune using time-aware cross-validation.
- Use time-aware cross-validation (e.g.,
TimeSeriesSplit). - Tune your classifier’s hyperparameters.
- Use a macro-averaged metric during tuning.
- Report the best parameters and the cross-validated performance.
Explain why your chosen metric is appropriate.
6.6 Evaluate classification performance
Now evaluate the tuned classifier on the held-out test set.
You should report:
- The classification report
- Multiclass ROC AUC
Then interpret:
- Which class is hardest to predict?
- Whether errors are symmetric or directional.
- Whether the model generalises plausibly forward in time.
Evaluate your tuned classifier on the test set and interpret the results.
6.6.1 Confusion matrix
A confusion matrix provides insight into the structure of classification errors.
Plot the confusion matrix and answer:
- Which misclassifications are most common?
- Are extreme classes confused with each other?
- What operational interpretation can you give to the error pattern?
- Are all mistakes equally costly in this operational setting?
6.6.2 Calibration of predicted probabilities
High ROC AUC does not guarantee that predicted probabilities are trustworthy.
Calibration checks whether predicted probabilities match observed frequencies.
- Plot calibration curves for each class.
- Carefully check the internal class order of your classifier before aligning probability columns.
- Interpret whether the classifier appears overconfident or underconfident.
- If probabilities were miscalibrated, how might that affect decision thresholds?
6.7 Exploring imbalance strategies
Even if the dataset appears balanced, use this as a controlled experiment.
Using the cross-sectional random CV design (not time-aware CV):
- Try oversampling and/or undersampling.
- Compare recall for the hardest class.
- Explain why these methods are not appropriate inside time-aware validation.
Oversampling/undersampling are mentioned to guide exploration — they are not appropriate inside TimeSeriesSplit because they would invalidate temporal ordering.
7 To read further about Pipelines
To read further about Pipelines’s functionalities, you could have a look at:
- this Medium post
- this tutorial on Python-Bloggers
- or look at the
Pipelinedocumentation and examples that come with it.
