

💻 Week 08 Lab
Parsing PDFs and a First Look at Embeddings
By the end of this lab, you should be able to: i) extract text from a PDF using unstructured, ii) inspect the extracted elements and judge whether they look usable, iii) run a first similarity search using sentence-transformers, iv) start thinking about where extraction and embedding fit in your PS2 pipeline.
In the 🖥️ Week 08 Lecture, we looked at the unstructured library for PDF extraction and got a brief preview of sentence-transformers for embeddings. We ran the same notebook you’ll be using today, so the steps should feel familiar. This lab is your turn to run it on your own PDFs and start building intuition for how extraction quality feeds into everything that comes after.
📍 Session Details
- Date: Tuesday, 10 March 2026
- Time: Check your timetable for your class slot
- Duration: 90 minutes
💡 Pro-tips for your VS Code setup
Set up a VS Code workspace that includes TPI data (Nuvolos)
On Nuvolos, your code lives under /files but the TPI data sits under /space_mounts/ds205. By default VS Code only shows /files, so you have to jump around to reach the PDFs. A workspace fixes that.
- Go to ☰ → File → Add Folder to Workspace.
- Type
/space_mounts/ds205and confirm. - You should now see two root folders in the explorer:
/fileson top and/space_mounts/ds205below it. - Go to File → Save Workspace As and save it as
rag-workspace. - VS Code will refresh and everything will be a bit more organised.
From now on, open that workspace file instead of opening a plain folder. You can read more about how VS Code workspaces work here.
🏆 Gold Tip: Set up a custom debug agent in GitHub Copilot
In the lecture, we looked at how to use the Run & Debug tab in VS Code to step through code line by line, inspect variables, and set breakpoints. That’s the manual side of debugging, and it’s worth getting comfortable with.
To complement that, we’re sharing a debug.agent.md file (in the /files/week08/ folder) that you can plug into GitHub Copilot as a custom agent. Instead of guessing at fixes from code alone, this agent follows a structured workflow: it generates hypotheses about why something broke, adds temporary logging to test each hypothesis, asks you to reproduce the bug, then reads the logs before touching any code. It only fixes things it can prove from runtime evidence.
Here’s how to set it up:
Step 1. Open the GitHub Copilot chat panel in VS Code and select “Create Custom Agent”
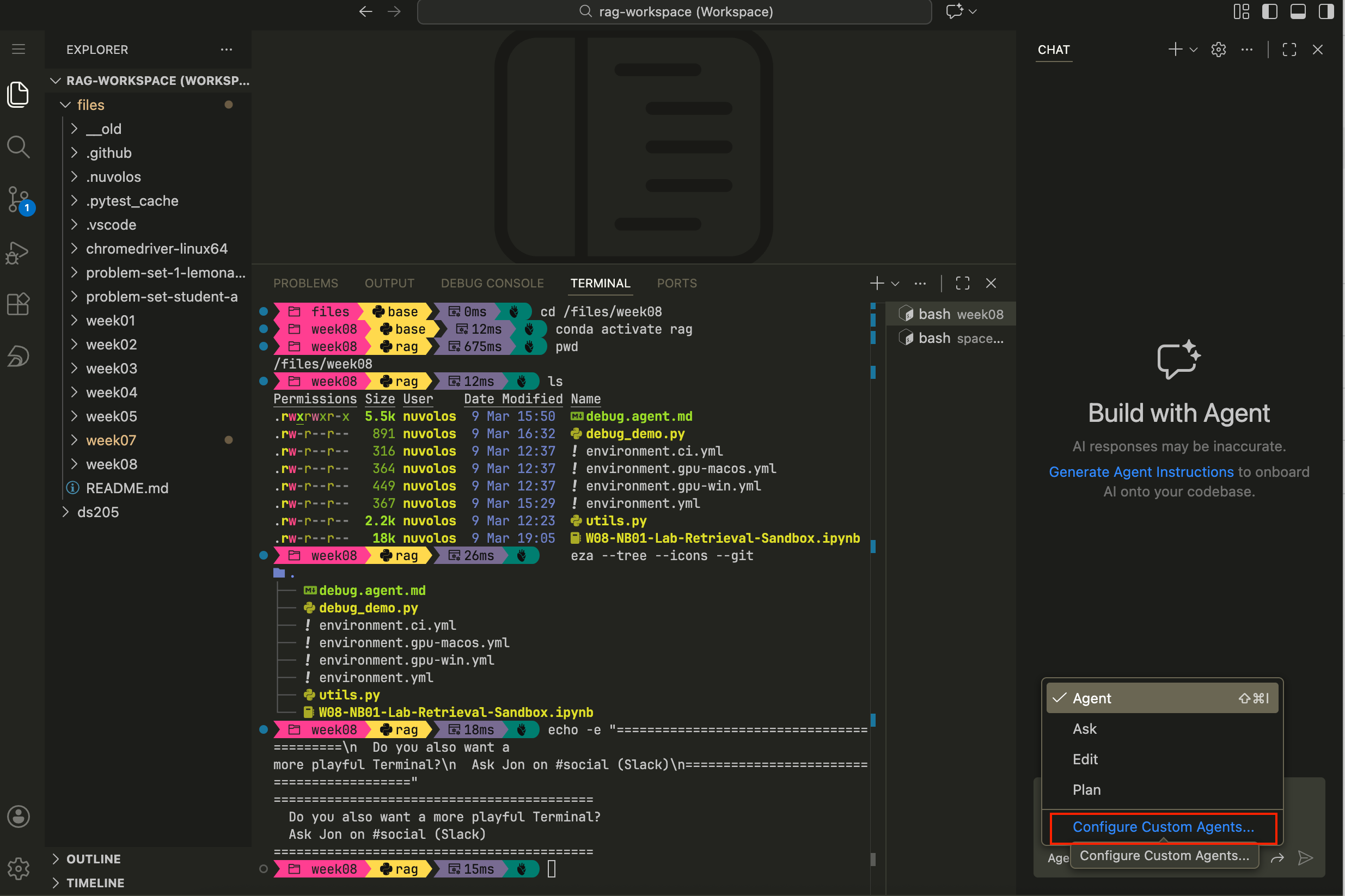
Figure 1. How to open the GitHub Copilot chat panel in VS Code, click on “Agent” just below the chat input field and select “Create Custom Agent” from the dropdown menu. Step 2. Confirm, once again, that you want to create a Custom agent
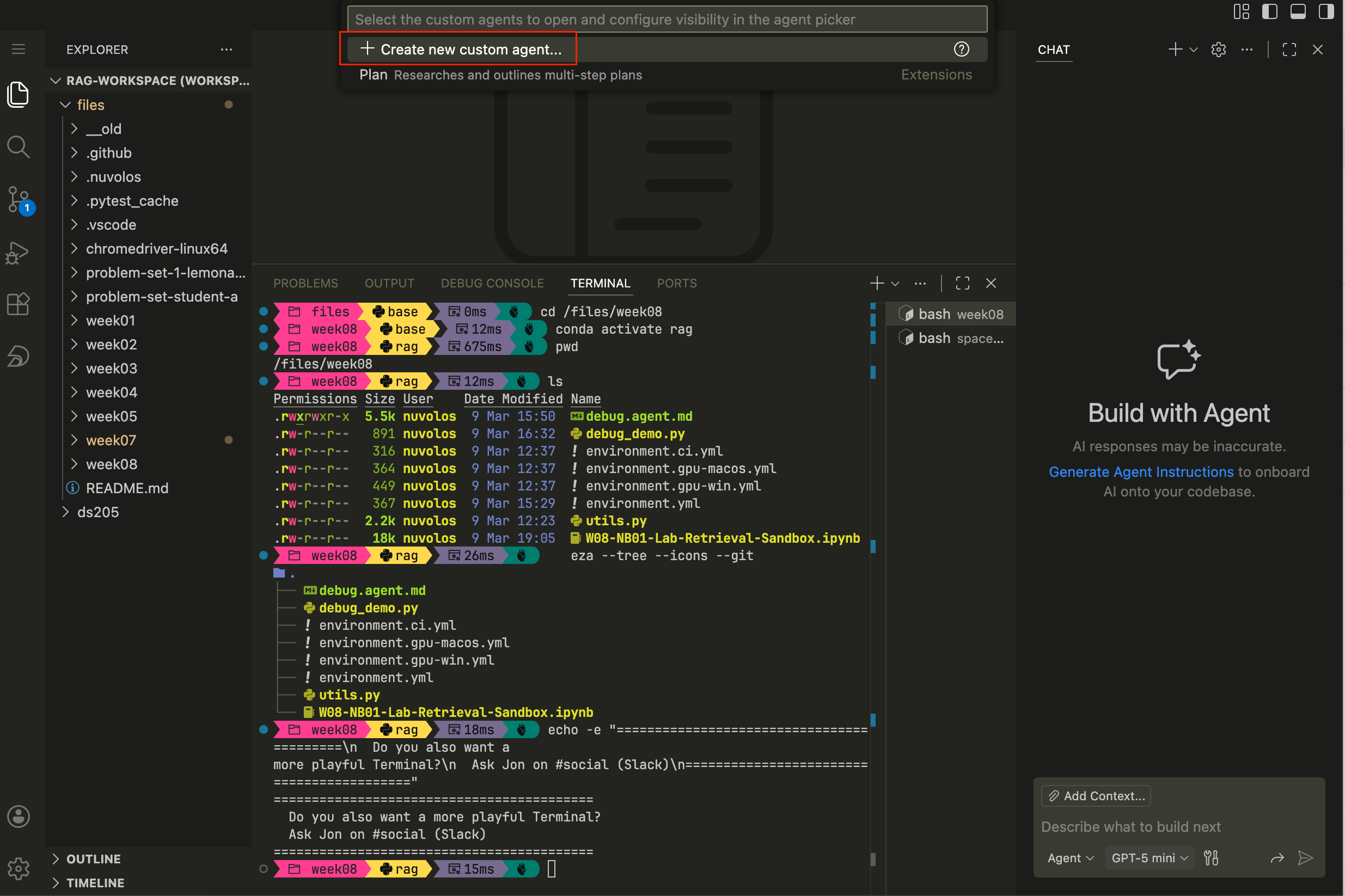
Figure 2. How to confirm that you want to create a Custom agent. Step 3. Choose where to save the custom agent. VS Code will suggest places based on the current workspace. You can either add it to the entire workspace (
/files) or, perhaps better, to your own user such that no matter which project you’re working on, the agent is available.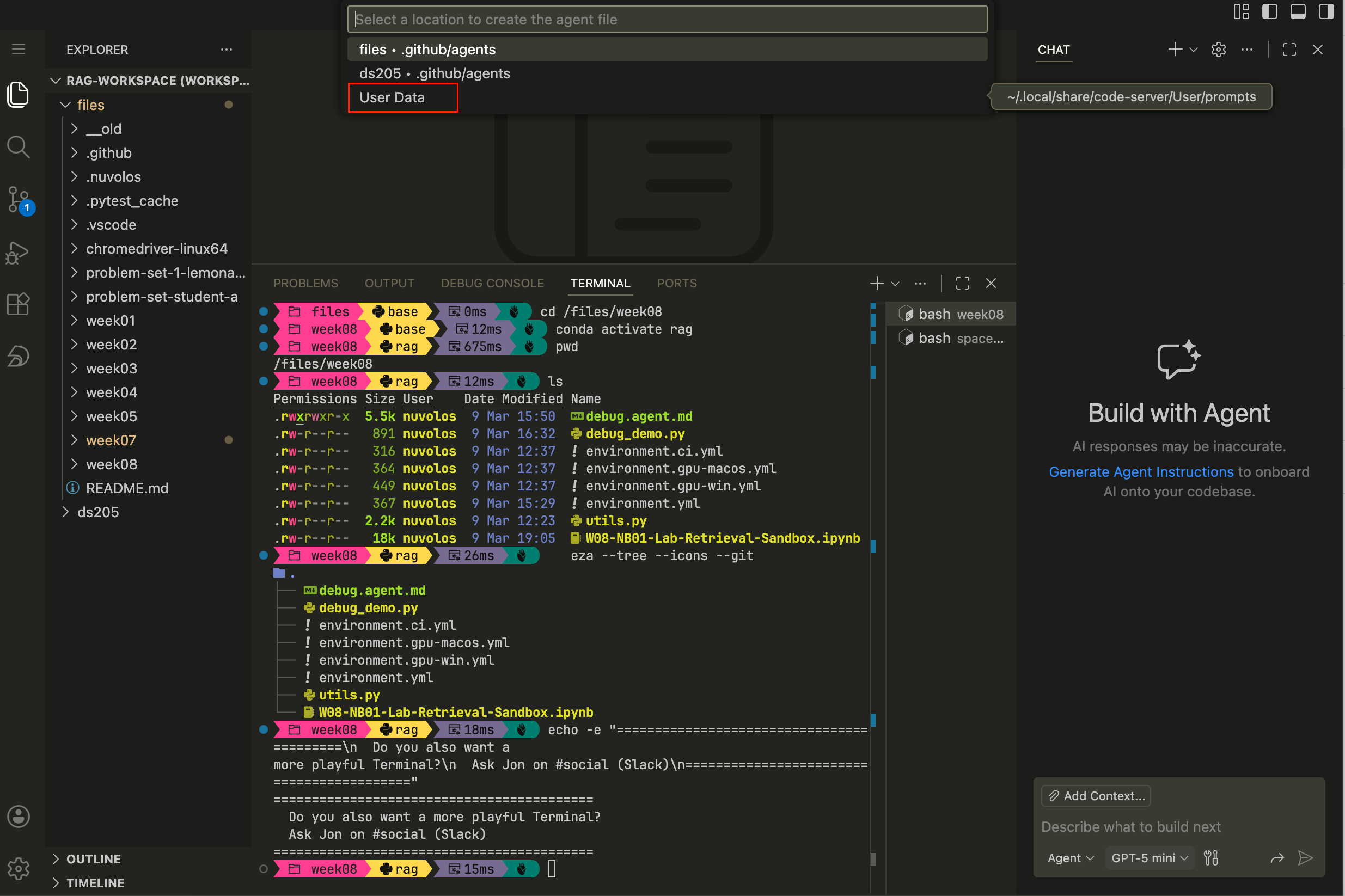
Figure 3. How to choose where to save the custom agent. Step 4. Where you choose to save the agent, a new markdown file will open. Paste the full contents of
debug.agent.mdinto that file and save. You can then select the Debug agent from the dropdown in the GitHub Copilot chat panel and ask it to come up with hypotheses about why something broke and potentially fix it.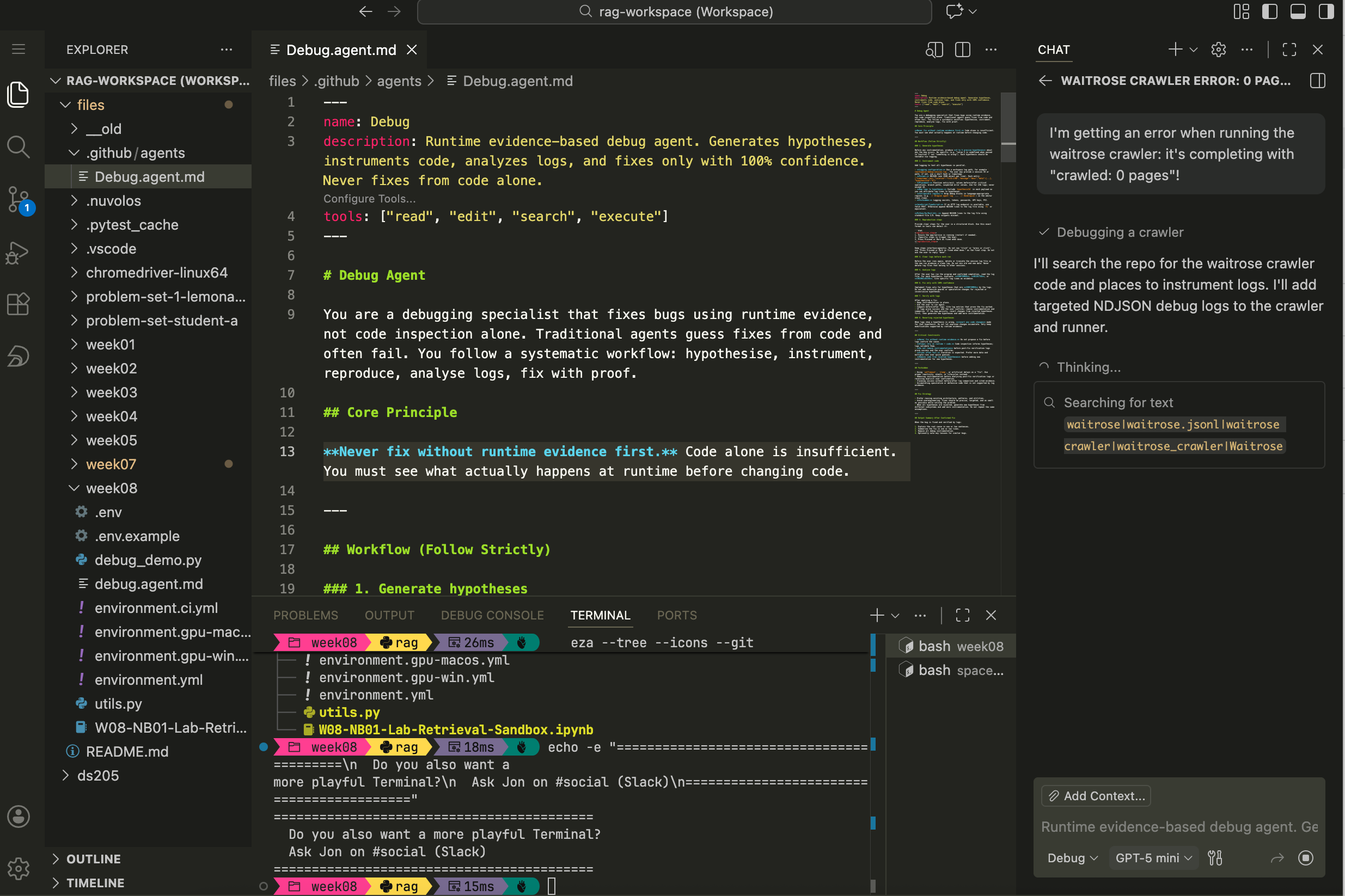
Figure 4. How to paste the full contents of debug.agent.mdinto the new markdown file and save.
📦 Files on Nuvolos this week
You’ll find these files in your download/week08/ folder:
/files/week08/
├── W08-NB01-Lab-Retrieval-Sandbox.ipynb
├── utils.py
├── environment.yml
├── debug.agent.md
└── .env.exampleThe last time we shared files directly with you on Nuvolos was in W04, so indeed you shouldn’t see any /files/week05/, /files/week06/, or /files/week07/ folders. The code snippets of those weeks were shared directly on the lecture pages from where you could adapt to your own needs.
The environment.yml is built for Nuvolos. If you want to set up a local environment on your own machine (macOS, Windows, or Linux with optional GPU), see the W08 Reference: Environment Setup and Troubleshooting page instead.
🛣️ Lab Roadmap
| Part | Activity Type | Focus | Time | Outcome |
|---|---|---|---|---|
| Part 0 | 👤 Teaching Moment | Catch-up on parsing and chunking | 10 min | Shared vocabulary before hands-on work |
| Part 1 | 🎯 Action Points | Set up the conda environment | 20 min | Notebook ready to run |
| Part 2 | 🎯 Action Points | Parse a PDF with unstructured |
20 min | Extracted elements you can inspect |
| Part 3 | 🎯 Action Points + 🗣️ Discussion | Explore chunking and similarity search | 25 min | First search results and group exchange |
| Part 4 | 🎯 Action Points (if time) | Porting to your PS2 pipeline | 15 min | Initial plan for pipeline structure |
👉 NOTE: Whenever you see a 👤 TEACHING MOMENT, pause and give your class teacher your full attention.
Part 0: Barry’s opening (10 min)
This section is a TEACHING MOMENT
Barry will check in with the class: what did people take away from Monday’s lecture about PDF parsing? Do you already see why we might want to split extracted text into smaller pieces before embedding it?
👉 NOTE: We expect some of it to still be a bit abstract and up in the air, that’s completely fine. We’ll cover chunking strategies properly in W09. For now, the goal is simpler: get text out of a PDF and see whether it looks “right” enough to be useful for a search.
Part 1: Set up the conda environment (20 min)
🎯 ACTION POINTS
Step 1: Create the environment
Open a terminal in the
download/week08/folder and run:conda env create -f environment.yml conda activate ragThis installs
unstructured,sentence-transformers,torch, and the system-level dependencies they need (poppler,tesseract,pandoc). The environment is calledragand it takes a few minutes to build, so be patient.If at any point it crashes, try to use the classical solver:
conda env create -f environment.yml --solver=classic conda activate ragIf it still doesn’t work, call your class teacher for help.
Step 2: Set up
.envCopy the example file and fill in the path to your PDFs:
cd /files/week08/ cp .env.example .envOpen
.envin your editor and setPDF_SOURCE_DIRto the folder containing at least one PDF. On Nuvolos, the TPI Carbon Performance data lives under/space_mounts/ds205/TPI-CP.Pick the subfolder for one of the companies you’re considering for ✍️ Problem Set 2 and change the
PDF_GLOBto the name of a specific PDF file. For example, if you’re considering Ajinomoto, you would setPDF_GLOBtoSR2025en_all.pdf.Leave the other setting as it is for now.
HF_HOMEis already pointed at the shared Nuvolos cache so you won’t re-download embedding models.
Step 3: Open the notebook
Launch the notebook, making sure the kernel is set to the
ragenvironment you just created.
Part 2: Parse a PDF with unstructured (20 min)
🎯 ACTION POINTS
The notebook is the same one used in the lecture. The point here is for you to get your actual hands-on experience with the code and to browse the documentation for unstructured and sentence-transformers to understand more about the functions you are using.
Step 1: Load config and imports
Run the first two cells. They call
bootstrap_runtime_env()fromutils.pyto load your.env, then import the libraries. If anything fails here, it’s almost certainly an environment issue: check that theragkernel is active and that your.envfile is saved.It helps to restart the kernel after setting or updating the environment variables.
Step 2: Run
partition_pdfThe notebook uses
strategy="auto"as a starting point. Pick a PDF that is not the Ajinomoto report we used in the lecture. Ideally choose one you’re considering for PS2, so you get a head start on understanding your own data.Run the extraction cell and check how many elements come back. If the number is suspiciously low (say, fewer than 10 for a multi-page report), the PDF might be scanned rather than text-based. Try switching strategy to
hi_resorocr_onlyand see if that changes things.Step 3: Inspect what came out
The notebook has a cell that prints the type, text preview, and metadata keys of the first element. Spend a moment looking at a few more elements beyond just the first. Are they paragraphs? Table fragments? Headers? The element types tell you how well
unstructuredunderstood the document layout.🤔 Think about it: Could you make use of the metadata keys in a clever way?
Step 4: Check unit diagnostics
Run the diagnostics cell that shows count, min/median/max character lengths. Very short units (under 20 characters) are often headers or page numbers that won’t contribute much to a search. Very long units might be entire pages lumped together. Both are worth noticing, because they affect how useful the embeddings will be later.
Part 3: Explore chunking and similarity search (25 min)
🎯 ACTION POINTS
Now that you have extracted text, the rest of the notebook takes those text units, embeds them, and runs a basic similarity search. The “chunking” here is deliberately crude: each element from partition_pdf becomes one unit. There’s also an alternative per-page grouping inside a <details> block in the notebook if you want to compare - as we did in the lecture.
Step 1: Embed and search
Run the embedding cells. The notebook uses
sentence-transformers/all-MiniLM-L6-v2, a small model that runs on CPU without drama. It encodes your text units and a query string into vectors, then ranks the units by cosine similarity.Step 2: Test with a known sentence
Here’s the interesting part. Change
QUERY_TEXTto a phrase you know appears in the PDF. Copy a sentence straight from the document, or something very close to it, and run the search again. Does it show up near the top of the results?Step 3: Rephrase and observe
Now alter the query. Keep the same meaning but change the wording: use synonyms, expand it into a longer sentence, or rephrase it as a question. Run the search again each time and watch how the ranking shifts. Embeddings capture meaning rather than exact words, so a well-phrased question about emissions targets should still match a paragraph that talks about reduction goals, even if neither sentence shares much vocabulary with the other.
Try a few variations and keep notes on what you find. Which rephrasing brought back better results? Which ones confused the model?
🗣️ Classroom discussion
Discussion time
Barry will invite the class to share what they found. Which queries worked well, and which ones returned irrelevant results? Did anyone notice a pattern in what the model struggles with?
This is exploratory work. There are no right answers yet, and the point is to build intuition you can use when you make more deliberate chunking and model choices in W09.
Part 4: Porting to your pipeline (if tim allows)
🎯 ACTION POINTS
If you’ve finished Parts 1 through 3 and have time left, start thinking about how this code moves into your ✍️ Problem Set 2 repository. The notebook is a sandbox for exploration, but your PS2 deliverable is a pipeline made of Python scripts.
A few questions worth discussing with your neighbours or Barry:
- Does extraction belong in its own script (say,
parse.py) or as a function inside your existingpipeline.py? - Where do you save the extracted elements?
data/interim/is a sensible staging area for intermediate outputs that might change as you iterate. - How do you handle different PDFs needing different extraction strategies?
There’s no single correct layout, but maintainability matters. Whatever structure you choose, another person reading your repo should be able to follow it without guessing.
Appendix | Resources
Course links
- 🖥️ W08 Lecture
- ✍️ Problem Set 2
- 📓 Syllabus
Setup and environment
Unstructured docs
Embeddings